Publications of the first funding period
Publications under funding of the Crc 1507
(sorted by year and alphabetical order of first author)
Asido M*, Lamm GHU, Lienert J, La Greca M, Kaur J, Mayer A, Glaubitz C, Heberle J, Schlesinger R, Kovalev K, Wachtveitl J* (2025) A detailed view on the (re)isomerization dynamics in microbial rhodopsins using complementary near-UV and IR readouts. Angew Chem Int Ed 64: e202416742
Berth SH, Vo L, Kwon DH, Grider T, Damayanti YS, Kosmanopoulos G, Fox A, Lau AR, Carr P, Donohue JK, Hoke M, Thomas S, Karam C, Fay AJ, Meltzer E, Crawford TO, Gaudet R, Shy ME, Hellmich UA, Lee SY, Sumner CJ, McCray BA* (2025) Combined clinical, structural and cellular studies discriminate pathogenic and benign TRPV4 variants. Brain 148(2): 564-579
Catapano C, Dietz MS, Kompa J, Jang S, Freund P, Johnsson K, Heilemann M* (2025) Long-term single-molecule tracking in living cells using weak-affinity protein labeling. Angew Chem Int Ed 64: e202413117
Hoffmann PC, Kim H, Obarska-Kosinska A, Kreysing JP, Andino-Frydman E, Cruz-León S, Margiotta E, Cernikova L, Kosinski J, Turoňová B, Hummer G*, Beck M* (2025) Nuclear pore permeability and fluid flow are modulated by its dilation state. Mol Cell 85, 1-18
Jackel E, Lazzeri G, Covino R* (2025) Free energy, rates, and mechanism of transmembrane dimerization in lipid bilayers from dynamically unbiased molecular dynamics simulations. J Phys Chem B 129: 1586-1596
Joseph B* (2025) Protein-protein interaction and conformational change in the alpha-helical membrane transporter BtuCD-F in the native cellular envelope. Chem Bio Chem 26: e202400858
Just A, Niemann N, Gnau P, Kock D, Heimerl T, Essen L-O, Batschauer A, Morgner N* (2025) Mechanistic insight into the oligomerization of Arabidopsis CRY1 and its inhibition by BIC1. bioRxiv, 2025.10.06.680677
Klausnitzer A, Kaur J, Rath T, Seidl S, Becker-Baldus J, Morgner N, Glaubitz C* (2025) Conformational Plasticity of LptC Regulates Lipopolysaccharide Transport by the LptB2FGC Complex. J Am Chem Soc 147: 35718
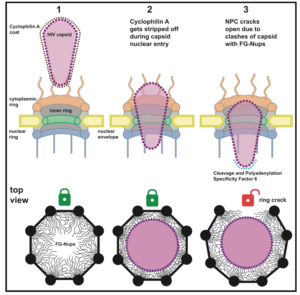
Kreysing JP, Heidari M, Zila V, Cruz-León S, Obarska-Kosinska A, Laketa V, Rohleder L, Welsch S, Köfinger J, Turoňová B, Hummer G*, Kräusslich HG*, Beck M* (2025) Passage of the HIV capsid cracks the nuclear pore. Cell 188, 930-943.e211
Krishnathas R, Mineev KS, Fourkiotis NK, Touret F, Sideras-Bisdekis C, Tsika AC, Gande SL, Linhard V, Sreeramulu S, Lennartz F, Weiss MS, Coutard B, Spyroulias GA, Schwalbe H* (2025) Structure-based rational design of a selective hydrolase inhibitor of the severe acute respiratory syndrome Coronavirus-2 Nsp3 macrodomain. Chem Bio Chem 2: e202500593
Lamm GHU, Zabelskii D, Balandin T, Gordeliy V, Wachtveitl J* (2025) Combined mutational and spectroscopic study on the calcium-related kinetic effects on the VirChR1 photocycle. J Phys Chem B 129: 2946-2957
Lasham J*, Djurabekova A, Kolypetris G, Zickermann V, Vonck J, Sharma V* (2025) Assessment of amino acid charge states based on cryo-electron microscopy and molecular dynamics simulations of respiratory complex I. Biochim Biophys Acta Bioenerg 1866: 149512
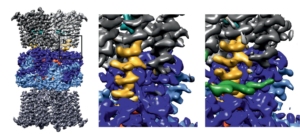
Lazarova M, Eicher T, Börnsen C, Zeng H, Athar M, Okada U, Yamashita E, Spannaus IM, Borgosch M, Cha HJ, Vargiu AV, Murakami S*, Diederichs K*, Frangakis AS*, Pos KM* (2025) Conformational plasticity across phylogenetic clusters of RND multidrug efflux pumps and its impact on substrate specificity. Nat Commun, accepted, bioRxiv2024/624703
Lazzeri G, Bolhuis PG, Covino R* (2025) Optimal rejection-free path sampling. Preprint at arXiv 2503.21037
Li Y, Niemann HH, Barth HD, Dietz MS, Heilemann M* (2025) Single-molecule FRET-tracking of InlB-activated MET receptor in living cells. bioRxiv 2025.05.06.652421
Mineev KS*, Gande SL, Wenzel A, Linhard V, Andrews-Clark H, Witt K, Köster S, Segarra M, McDowell MA, Acker-Palmer A, Schwalbe H* (2025) Structural role of conformational heterogeneity and juxtamembrane regions in the Ephrin A1 interactions with the EphA2 receptor ligand-binding domain. J Mol Biol, in press
Mineev KS, Gande SL, Linhard V, Moghaddam SK, Schwalbe H* (2025) NMR resonance assignment of a ligand-binding domain of ephrin receptor A2. Biomol NMR Assign 19: 23-28
Pecorilla C, Altmeyer A, Haapanen O, Lee Y, Zickermann V*, Sharma V* (2025) Conformational dynamics of a histidine molecular switch in a cation/proton antiporter. Biochim Biophys Acta Bioenerg 1866: 149563
Schulte J, Schwegler E, Hellmich UA, Morgner N* (2025) Quantification of protein homodimer affinity using native mass spectrometry. arXiv, 2510.08324
Spinetti E, Karagöz GE, Covino R* (2025) The structural dynamics of IRE1 and its interaction with unfolded peptides.eLife 14: RP106716
Stautz J, Griwatz D, Kaltwasser S, Mehdipour AR, Ketter S, Thiel C, Wunnicke D, Schrecker M, Mills DJ, Hummer G, Vonck J*, Hänelt I* (2025) A short intrinsically disordered region at KtrB’s N-terminus facilitates allosteric regulation of K+ channel KtrAB. Nat Commun 16: 4252
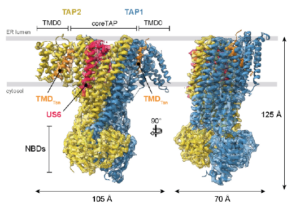
Stolz M, Sušac L, Fahim A, Keller R, Saggau L, Mancia F, Trowitzsch S*, Tampé R* (2025) Architectural principles of transporter-chaperone coupling within the native MHC I peptide-loading complex. Science Adv, accepted
Taniguchi R, Orniacki C, Kreysing JP, Zila V, Zimmerli CE, Bohm S, Turonova B, Kräusslich HG, Doye V*, Beck M* (2025) Nuclear pores safeguard the integrity of the nuclear envelope. Nat Cell Biol 27: 762-775
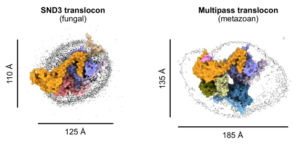
Yang TJ, Mukherjee S, Langer JD, Hummer G, McDowell MA* (2025) SND3 is the membrane insertase within a distinct SEC61 translocon complex. Nat Commun 16: 9566
Zabelskii D, Bukhdruker S, Bukhalovich S, Tsybrov F, Lamm GHU, Astashkin R, Doroginin D, Matveev G, Sudarev V, Kuzmin A, Zinovev E, Vlasova A, Ryzhykau Y, Ilyinsky N, Gushchin I, Bourenkov G, Alekseev A, Round A, Wachtveitl J, Bamberg E*, Gordeliy V* (2025) Ion-conducting and gating molecular mechanisms of channelrhodopsins revealed by true-atomic-resolution structures of open and closed states. Nat Struct Mol Biol 32: 1347-1357
Abraham BG, Haikarainen T, Vuorio J, Girych M, Virtanen AT, Kurttila A, Karathanasis C, Heilemann M, Sharma V, Vattulainen I, Silvennoinen O (2024) Molecular basis of JAK2 activation in erythropoietin receptor and pathogenic JAK2 signaling. Sci Adv 10: eadl2097
Betsennaia E, Maslov I, Balandin T, Alekseev A, Yudenko A, Abu Shamseye A, Zabelskii D, Baumann A, Catapano C, Karathanasis C, Gordeliy V, Heilemann M, Gensch T*, Borshchevskiy V* (2024) Revealing the oligomerization of channelrhodopsin-2 in the cell membrane using photo-activated localization microscopy. Angew Chem Int Ed 63: e202307555
Brunnberg J, Barends M, Frühschulz S, Winter C, Battin C, de Wet B, Cole DK, Steinberger P, Tampé R* (2024) Dual role of the peptide loading complex as proofreader and limiter of MHC-I presentation. Proc Natl Acad Sci USA 121: 22321600121
Dajka M, Rath T, Morgner N, Joseph B* (2024) Dynamic basis of lipopolysaccharide export by LptB2FGC. eLife, 2024.05.20.594984
Djurabekova A, Lasham J, Zdorevskyi O, Zickermann V, Sharma V* (2024) Long-range electron proton coupling in respiratory complex I – insights from molecular simulations of the quinone chamber and antiporter-like subunits.Biochem J 481: 499-514
Gopinath A, Rath T, Morgner N, Joseph B* (2024) Lateral gating mechanism and plasticity of the β-barrel assembly machinery complex in micelles and Escherichia coli. PNAS Nexus 3, 19
Johnson A G, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ* (2024) Structure and assembly of a bacterial gasdermin pore. Nature 628: 657-663
Kessler LF, Balakrishnan A, Menche T, Wang D, Li Y, Mantel M, Glogger M, Dietz MS, Heilemann M* (2024) Self-quenched fluorophore-DNA labels for super-resolution fluorescence microscopy. J Phys Chem B 128: 6751-6759
Kettel P, Marosits L, Spinetti E, Rechberger M, Giannini C, Radler P, Niedermoser I, Fischer I, Versteeg G A, Loose M, Covino R, Karagoz GE* (2024) Disordered regions in the IRE1α ER lumenal domain mediate its stress-induced clustering. EMBO J 43: 4668–4698
Lamm GHU, Zabelskii D, Balandin T, Gordeliy V, Wachtveitl J* (2024) Calcium-sensitive microbial rhodopsin VirChR1: A femtosecond to second photocycle study. J Phys Chem Lett 15: 5510-5516
Laube E, Schiller J, Zickermann V*, Vonck J* (2024) Using cryo-EM to understand the assembly pathway of respiratory complex I. Acta Crystallogr D 80: 159-173
Li Y, Arghittu SM, Dietz MS, Hella GJ, Haße D, Ferraris DM, Freund P, Barth H, Iamele L, de Jonge H, Niemann HH, Covino R*, Heilemann M* (2024) Single-molecule imaging and molecular dynamics simulations reveal early activation of the MET receptor in cells. Nat Commun 15: 9486
Llaó-Cid C*, Peguera B*, Kobialka P, Decker L, Vogenstahl J, Alivodej N, Srivastava S, Jin J, Kirchmaier BC, Milla C, Schlierbach H, Schänzer A, Acker T, Segarra M*, Acker-Palmer A* (2024) Vascular FLRT2 regulates venous-mediated angiogenic expansion and CNS barriergenesis. Nat Commun 15: 10372
Löffler M, Frühschulz S, Rockel Z, Pečak M, Tampé R*, Wieneke R* (2024) Antigen delivery controlled by an on-demand photorelease. Angew Chem Int Ed 63: e202405035
Mineev KS, Hargittay B, Jin J, Catapano C, Dietz MS, Segarra M, Harwardt MS, Richter C, Jonker HRA, Saxena K, Sreeramulu S, Heilemann M, Acker-Palmer A, Schwalbe H* (2024) Differential effects of the N-terminal helix of FGF8b on the activity of a small-molecule FGFR inhibitor in cell culture and for the extracellular domain of FGFR3c in solution. FEBS Lett 598: 2518-2532
Novischi SYP, Karoly-Lakatos A, Chok K, Bonifer C, Becker-Baldus J, Glaubitz C* (2024) Probing the allosteric NBD-TMD crosstalk in the ABC transporter MsbA by solid-state NMR. Commun Biol 7: 43
Podoliak E, Lamm GHU, Marin E, Schellbach AV, Fedotov DA, Stetsenko A, Asido M, Maliar N, Bourenkov G, Balandin T, Baeken C, Astashkin R, Schneider TR, Bateman A, Wachtveitl J, Schapiro I, Busskamp V, Guskov A, Gordeliy V, Alekseev A*, Kovalev K* (2024) A subgroup of light-driven sodium pumps with an additional Schiff base counterion, Nat Commun 15: 3119
Rentsch D, Bergs A, Shao J, Elvers N, Ruse C, Seidenthal M, Aoki I, Gottschalk A* (2024) Tools and methods for cell ablation and cell inhibition in Caenorhabditis elegans. Genetics 229: iyae119
Schwickert KK, Glitscher M, Bender D, Benz NI, Murra R, Schwickert K, Pfalzgraf S, Schirmeister T, Hellmich UA*, Hildt E* (2024) Zika virus replication is impaired by a selective agonist of the TRPML2 ion channel. Antiviral Res 228: 105940
Albertazzi L, Heilemann M* (2023) When weak is strong: A plea for low-affinity binders for optical microscopy. Angew Chem Int Ed 62: e202303390
Barends M, Koller N, Schölz C, Durán V, Bošnjak B, Becker J, Döring M, Blees H, Förster R, Kalinke U, Tampé R* (2023) Dynamic interactome of the MHC I peptide loading complex in human dendritic cells. Proc Natl Acad Sci USA120: e2219790120
Catapano C, Rahm JV, Omer M, Teodori L, Kjems J, Dietz MS, Heilemann M* (2023) Biased activation of the receptor tyrosine kinase HER2. Cell Mol Life Sci 80: 158.
Goretzki B, Wiedemann C, McCray BA, Schäfer SL, Jansen J, Tebbe F, Mitrovic SA, Nöth J, Cabezudo AC, Donohue JK, Jeffries CM, Steinchen W, Stengel F, Sumner CJ, Hummer G, Hellmich UA* (2023) Crosstalk between regulatory elements in disordered TRPV4 N-terminus modulates lipid-dependent channel activity. Nat Commun 14: 4165
Hargittay B, Mineev KS, Richter C, Sreeramulu S, Jonker HRA, Saxena K, Schwalbe H* (2023) NMR resonance assignment of a fibroblast growth factor 8 splicing isoform b. Biomol NMR Assign 17: 135-142
Jung H#, Covino R#, Arjun A, Leitlold C, Dellago C, Bolhuis P, Hummer G* (2023) Machine-guided path sampling to discover mechanisms of molecular self-organization. Nat Comput Sci 3: 334–345
Kessler LF, Balakrishnan A, Deußner-Helfmann NS, Li Y, Mantel M, Glogger M, Barth HD, Dietz MS, Heilemann M* (2023) Self-quenched fluorophore dimers for DNA-PAINT and STED microscopy. Angew Chem Int Ed 62: e202307538
Ketter S, Joseph B* (2023) Gd(III)-trityl-nitroxide triple labelling and distance measurements in the heterooligomeric cobalamin transport complex in the native lipid bilayers. J Am Chem Soc 145: 960–966
Lasham J*, Haapanen O, Zickermann V, Sharma V* (2023) Tunnel dynamics of quinone derivatives and its coupling to protein conformational rearrangements in respiratory complex I. Biochim Biophys Acta Bioenerg 1864: 148951
Lazzeri G, Jung H, Bolhuis PG, Covino R* (2023) Molecular free energies, rates, and mechanisms from data-efficient path sampling simulations. J Chem Theory Comput 19: 9060–9076
Rudolph M, Tampé R*, Joseph B* (2023) Time-resolved Mn2+–NO and NO–NO distance measurements reveal that catalytic asymmetry regulates alternating access in an ABC transporter. Angew Chem Int Ed 62: e2023070
Sagert L, Winter C, Ruppert I, Zehetmaier M, Thomas C, Tampé R* (2023) The ER folding sensor UGGT1 acts on TAPBPR-chaperoned peptide-free MHC I. eLife 12: e85432
Tröster A, Jores N, Mineev KS, Sreeramulu S, DiPrima M, Tosato G, Schwalbe H* (2023) Targeting EPHA2 with Kinase Inhibitors in Colorectal Cancer. Chem Med Chem 18: e202300420
Tröster A, DiPrima M, Jores N, Kudlinzki D, Sreeramulu S, Gande SL, Linhard V, Ludig D, Schug A, Saxena K, Reinecke M, Heinzlmeir S, Leisegang MS, Wollenhaupt J, Lennartz F, Weiss MS, Kuster B, Tosato G*, Schwalbe H* (2023) Optimization of the Lead Compound NVP-BHG712 as a Colorectal Cancer Inhibitor. Chemistry 29: e202203967
Yu M, Heidari M, Mikhaleva S, Tan PS, Mingu S, Ruan H, Reinkemeier CD, Obarska-Kosinska A, Siggel M, Beck M, Hummer G*, Lemke EA* (2023) Visualising the disordered nuclear transport machinery in situ. Nature 617: 162-169
Domnick A, Winter C, Sušac L, Hennecke L, Hensen M, Zitzmann N, Trowitzsch S, Thomas C, Tampé R* (2022) Molecular basis of MHC I quality control in the peptide loading complex. Nat Commun 13: 4701
Koller N, Höllthaler P, Barends M, Döring M, Spahn C, Duran V, Costa B, Becker J, Heilemann M, Kalinke U*, Tampé R* (2022) Nanoscale organization of the MHC I peptide loading complex in human dendritic cells. Cell Mol Life Sci 79: 477
Müller IK, Winter C, Thomas C, Spaapen RM, Trowitzsch S*, Tampé R* (2022) Structure of an MHC I-tapasin-ERp57 editing complex defines chaperone promiscuity. Nat Commun 13: 5283
Plé C, Tam HK, Vieira Da Cruz A, Compagne N, Jiménez-Castellanos JC, Müller RT, Pradel E, Foong WE, Malloci G, Ballée A, Kirchner MA, Moshfegh P, Herledan A, Herrmann A, Deprez B, Willand N, Vargiu AV, Pos KM*, Flipo M*, Hartkoorn RC* (2022) Pyridylpiperazine-based allosteric inhibitors of RND-type multidrug efflux pumps. Nat Commun13: 115
Schaefer SL, Hummer G* (2022) Sublytic Gasdermin-D pores captured in atomistic molecular simulations. eLife 11: e81432
Schiller J, Laube E, Wittig I, Kühlbrandt W, Vonck J*, Zickermann V* (2022) Insights into complex I assembly: Function of NDUFAF1 and a link with cardiolipin remodeling. Sci Adv 8: eadd3855
Steinhilper R, Höff G, Heider J, Murphy BJ* (2022) Structure of the membrane-bound formate hydrogenlyase complex from Escherichia coli. Nat Commun 13: 5395
Sušac L, Yuong MT, Thomas C, von Bülow S, O’Brien-Ball C, Santos AM, Fernandes RA, Hummer G, Tampé R*, Davis SJ* (2022) Structure of a fully assembled tumor-specific T-cell receptor ligated by pMHC. Cell 185: 3201-13
Winter C, Domnick A, Cernova D, Tampé R* (2022) Semisynthetic viral inhibitor for light control of the MHC I peptide loading complex. Angew Chem Int Ed 61: e202211826
Zhang L, Simonsen C, Zimova L, Wang K, Moparthi L, Gaudet R, Ekoff M, Nilsson G, Hellmich UA, Vlachova V, Gourdon P*, Zygmunt PM* (2022) Cannabinoid non-cannabidiol site modulation of TRPV2 structure and function. Nat Commun 13: 7483
*corresponding author(s) #equal contribution
Recent Highlights
Kreysing JP, Heidari M, Zila V, Cruz-León S, Obarska-Kosinska A, Laketa V, Rohleder L, Welsch S, Köfinger J, Turoňová B, Hummer G*, Kräusslich HG*, Beck M* (2025) Passage of the HIV capsid cracks the nuclear pore. Cell 188, 930-943.e211
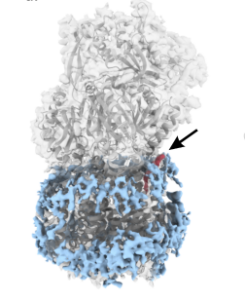
Lazarova M, Eicher T, Börnsen C, Zeng H, Athar M, Okada U, Yamashita E, Spannaus IM, Borgosch M, Cha HJ, Vargiu AV, Murakami S*, Diederichs K*, Frangakis AS*, Pos KM* (2025) Conformational plasticity across phylogenetic clusters of RND multidrug efflux pumps and its impact on substrate specificity. Nat Commun, accepted, bioRxiv 2024/624703
Stautz J, Griwatz D, Kaltwasser S, Mehdipour AR, Ketter S, Thiel C, Wunnicke D, Schrecker M, Mills DJ, Hummer G, Vonck J*, Hänelt I* (2025) A short intrinsically disordered region at KtrB’s N-terminus facilitates allosteric regulation of K+channel KtrAB. Nat Commun 16: 4252
Stolz M, Sušac L, Fahim A, Keller R, Saggau L, Mancia F, Trowitzsch S*, Tampé R* (2025) Architectural principles of transporter-chaperone coupling within the native MHC I peptide-loading complex. Science Adv, accepted
Yang TJ, Mukherjee S, Langer JD, Hummer G, McDowell MA* (2025) SND3 is the membrane insertase within a distinct SEC61 translocon complex. Nat Commun 16: 9566